Research News
May 14, 2026
- Medicine
Two proteins drive fibrosis — Scientists show they can be blocked
How immune cells drive liver scarring
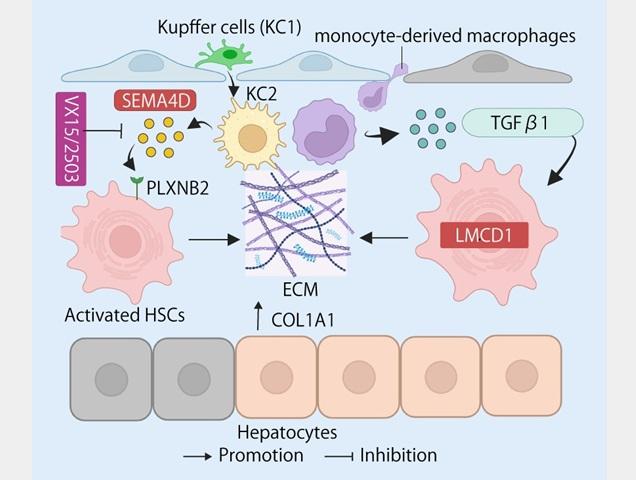
Various liver cell types interact to drive fibrosis during chronic liver disease. Kupffer cells (KC1) undergo phenotypic changes, transitioning to an activated state (KC2), accompanied by the accumulation of monocyte-derived macrophages. These macrophages promote hepatic stellate cell (HSC) activation through two distinct signaling pathways. One pathway operates via TGF-β1 and the transcription factor LMCD1, keeping HSCs locked in a fibrogenic state. A second pathway involves SEMA4D binding to its receptor PLXNB2 on HSCs. Blocking SEMA4D with an experimental antibody (VX15/2503) disrupts this signaling, reducing collagen production and scar formation.
Credit: Osaka Metropolitan University
The liver is often called a “silent organ,” as it can sustain significant damage without obvious symptoms. But when injury is prolonged, whether from alcohol, poor diet, or chronic hepatitis virus infection, it triggers fibrosis: a progressive hardening and scarring of liver tissue.
Without effective intervention, fibrosis can advance to cirrhosis and liver cancer, one of the leading causes of cancer death worldwide. Despite decades of research, no approved therapy currently exists that can reliably halt or reverse fibrosis once it is established.
What makes fibrosis particularly difficult to treat is that it does not behave uniformly. Some patients, such as those who successfully achieve viral clearance of hepatitis C, show spontaneous regression of fibrosis, whereas others continue to deteriorate. Understanding why the liver can heal in some cases but not others has been a central and largely unanswered question in research.
A team from Osaka Metropolitan University, led by Associate Professor Le Thi Thanh Thuy and Professor Norifumi Kawada used a cutting-edge technique called single-cell fixed RNA profiling (FLEX)—applied here to liver fibrosis research for the very first time—to analyze the gene activity of approximately 38,000 individual liver cells from healthy, fibrotic, and recovering mouse livers.
FLEX produces a precise picture of each individual cell’s behavior, even from frozen tissue samples. The result is a comprehensive cellular “atlas”—a detailed map of how every major liver cell type changes as fibrosis progresses and, critically, as it retreats.
Using the atlas, the researchers showed that hepatocytes in the pericentral zone, the central region of each liver lobule, show a striking restoration of their normal gene activity during recovery. This suggests that this specific hepatocyte population plays an active role in driving liver regeneration, offering a new direction for understanding how the organ heals itself.
The atlas revealed two molecular drivers of fibrosis that stood out as therapeutic targets.
The first was a protein called semaphorin-4D (SEMA4D). During fibrosis, immune cells called monocyte-derived macrophages secrete SEMA4D. The protein acts as a “distress signal,” binding to a receptor called Plexin B2 on hepatic stellate cells. This binding activates stellate cells and ramps up collagen production, worsening the scarring.
When the research team treated fibrotic mice with VX15/2503, a humanized monoclonal antibody that blocks SEMA4D, fibrosis was significantly reduced.
Crucially, elevated SEMA4D was also detected in human liver biopsies from patients with advanced fibrosis, and its levels declined in patients who cleared hepatitis C, a decline that correlated with a reduced risk of liver cancer progression.
The second target was LIM and cysteine-rich domains 1 (LMCD1), a transcription factor that operates inside hepatic stellate cells themselves.
Rather than receiving a signal from outside, LMCD1 acts as an internal “master switch” that keeps stellate cells locked in an active, scar-producing state. It was found to be highly expressed during fibrosis and suppressed during regression.
Silencing LMCD1 in the laboratory reduced fibrotic protein production, whereas forcing its overexpression had the opposite effect, driving fibrosis through the AKT/mTOR signaling pathway.
Like SEMA4D, LMCD1 levels in human liver samples correlated with fibrosis severity across both fatty liver disease and hepatitis C patient cohorts.
The evidence suggested that these two molecules were driving fibrosis in human patients and, most importantly, could be blocked. By targeting both proteins using a combination therapy, patients could potentially reduce the progression of the disease.
“What excites us most is that SEMA4D and LMCD1 work through completely different mechanisms and in two different cell types—macrophages and stellate cells,” Associate Professor Le Thi Thanh Thuy, the co-corresponding author of the study said. “When we analyzed human liver biopsy samples, patients with advanced fibrosis had high levels of both proteins, and both declined when the disease improved. This confirms these targets are relevant to human disease, not just mice.”
The difference between patients who heal and those who do not may lie in whether their livers can successfully shut down these pro-fibrotic signals. By identifying the precise proteins and cell interactions that tip this balance, the team hopes their work will inform the development of the first genuinely effective antifibrotic therapies.
“Finding these two targets points toward a combination antifibrotic therapy that attacks fibrosis from multiple angles simultaneously,” Professor Thuy continued. “The goal is not just to slow scarring down, but to actively stop it and push the liver toward recovery, which could meaningfully reduce the risk of liver cancer and lift the enormous burden this disease places on patients.”
The study was published in JHEP Reports.
Funding
PMD was awarded the Japanese Government (MEXT) Scholarship for the Ph.D. course.
LTTT received a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (JSPS Kaken C, 22K08083).
NK received a Grant-in-Aid for Scientific Research from JSPS (J192640002), and a grant from the Japan Agency for Medical Research and Development (AMED-24fk0210107h0003).
Paper information
Journal: JHEP Reports
Title: Single-cell fixed RNA profiling uncovers SEMA4D and LMCD1 as therapeutic targets in a liver fibrosis model
DOI: 10.1016/j.jhepr.2026.101783
Authors: Pham Minh Duc, Le Thi Thanh Thuy, Hoang Hai, Tran Van Bao, Nguyen Thi Ha, Pham Tuan Anh, Hiroko Ikenaga, Hideki Fujii, Kanako Yoshida, Masaru Enomoto, Sawako Uchida-Kobayashi, Hideto Yuasa, Tsutomu Matsubara, Vu Thuong Huyen, Yi Cheng, Ryota Yamagishi, Naoko Ohtani, Daisuke Oikawa, Fuminori Tokunaga, Tatiana Kisseleva, David A. Brenner, Yasuko Iwakiri, Jordi Gracia-Sancho, and Norifumi Kawada
Published: 11 February 2026
URL: https://doi.org/10.1016/j.jhepr.2026.101783
Contact
Le Thi Thanh Thuy
Graduate School of Medicine
Email: thuy[at]omu.ac.jp
*Please change [at] to @.
SDGs
